:focal(smart))
How notebook.link Scales Without Backends
This is a guest blog post by Sylvain Corlay, CEO of QuantStack. QuantStack has recently launched notebook.link - a platform to run computational notebooks in WebAssembly on top of prefix' infrastructure.
Traditional Jupyter deployments rely on backend infrastructure to execute code and manage kernels. For this reason, scaling computational notebooks has always been a trade-off: more users mean more servers, more maintenance, and more cost. What if you could eliminate the backend entirely?
notebook.link does that! It combines WebAssembly and the conda ecosystem to deliver scalable, serverless computing environments that run entirely in the browser. Without a backend there are no scaling headaches and we can offer instant, collaborative scientific computing for everyone. Key technologies underlying this product are
the prefix.dev service, which hosts the emscripten-forge package distributions,
and rattler, the high-performance dependency solver.
Together, they power notebook.link’s ability to deliver full conda environments in the browser, without the need for backend compute to scale with user demand. The modernised conda stack introduced by prefix.dev was instrumental in making this project possible, providing the tools and infrastructure needed to build, resolve, and distribute WebAssembly-based scientific computing environments at scale.
Scaling Jupyter
Most platforms offering scientific computing environments face a fundamental scalability issue: backend resources must grow proportionally with usage. Each new user or session typically requires additional server capacity, leading to:
Higher infrastructure costs as user bases expand.
Performance bottlenecks during peak usage.
Complex maintenance to manage and scale backend resources.
For institutions and organizations, this model is unsustainable when serving large, global audiences or supporting collaborative workflows. For example, UC Berkeley’s “Data8” class serves 10,000 students and requires over $100,000 in cloud resources annually, along with dedicated staff for operational maintenance.
What if you could support hundreds of thousands of users - without scaling a single server?
JupyterLite to the rescue
JupyterLite is a Jupyter distribution that runs entirely in the browser using WebAssembly. Unlike traditional Jupyter deployments, it operates as a static website, requiring no server-side infrastructure.
Static Deployment: Served as static files, JupyterLite eliminates the need for backend compute. Deployments like https://jupyter.org/try leverage this approach to serve 400,000 unique visitors monthly.
Instant Access: Users launch a full JupyterLab environment directly in their browser- no installation or configuration needed.
Scalability: Without servers to maintain, scaling is effortless, even as usage grows.
JupyterLite’s development began in 2021 at QuantStack. Beyond contributions from QuantStack, it has received significant contributions from a vibrant community and has achieved widespread adoption, powering interactive computing environments at scale across the web. In early 2026, JupyterLite officially joined the Jupyter Project, transitioning from a company-backed initiative to a recognized standard in the ecosystem.
Yet, the availability and flexibility of software packages remain pivotal to its continued success. This is where emscripten-forge comes in.
Emscripten-forge: a Unified Software Distribution for the Web
Building a new software distribution from the ground up is a monumental task, especially when targeting a new platform. With emscripten-forge, we didn’t port a few Python packages to the web. We set ourselves up to recreating the entire conda ecosystem in the browser, enabling users to run Python, R, C++, GNU Octave, including legacy Fortran-based library, and console applications, natively, without installation or backend servers.
This means:
Thousands of packages: From NumPy and SciPy to R’s tidyverse and Fortran-based HPC libraries, emscripten-forge provides the tools researchers, educators, and developers already use.
No compromises: Users can install packages with mamba, just like on their local machines - except now, it all runs in the browser.
Cross-language support: Thanks to the xeus kernel ecosystem, notebook.link supports Python, R, C++, and GNU Octave, all within the same environment.
Such a distribution had to be built upon solid foundations. This is where prefix.dev’s modernized conda stack came into play:
The Role of prefix.dev’s Modern Conda Stack
The success of emscripten-forge and notebook.link is deeply tied to the modern and consolidated conda stack developed by prefix.dev:
Rattler-build: Emscripten-forge uses rattler-build instead of conda-build. This allowed us to target the emscripten-wasm32 platform, which was not supported by conda-build. Rattler-build’s flexibility and modern design made it possible to compile complex scientific packages for WebAssembly.
The conda recipe format: The new recipe format introduced by prefix.dev, and proposed formally in a Conda Enhancement Proposal (CEP), was adopted by emscripten-forge from day one.
In-Browser Dependency Resolution: We use a WebAssembly build of rattler for dependency resolution directly in the browser. This enables real-time environment creation and package installation without server-side computation.
Hosting Infrastructure: The emscripten-forge package repository is hosted on prefix.dev’s servers, ensuring reliable, scalable, and fast global delivery of WebAssembly packages.
Without these innovations, building and distributing a full conda-based scientific computing environment in the browser would not have been feasible.
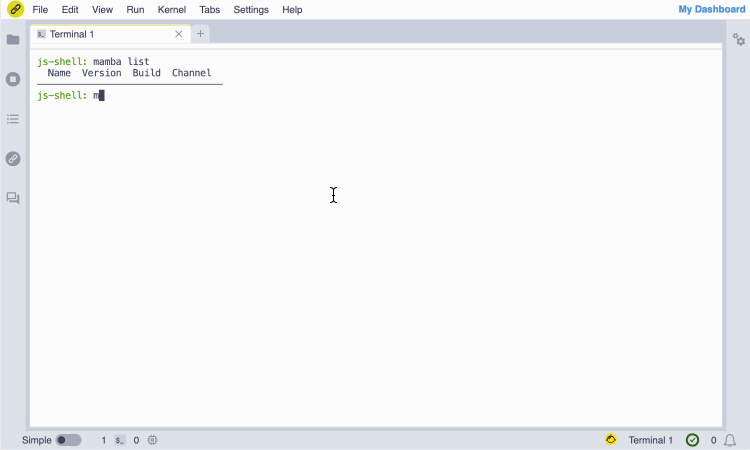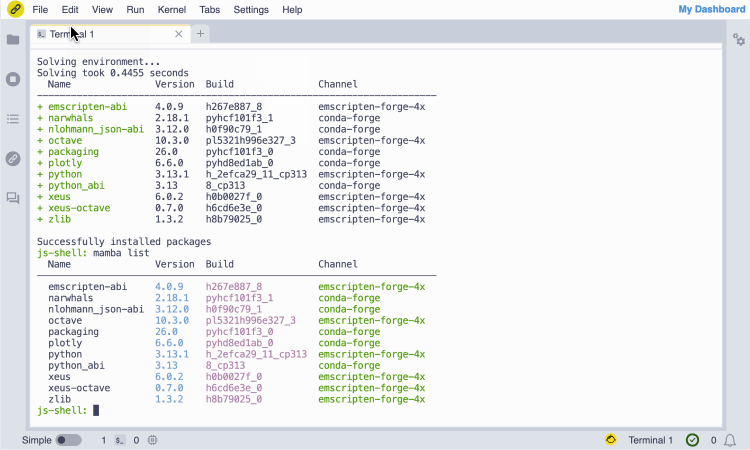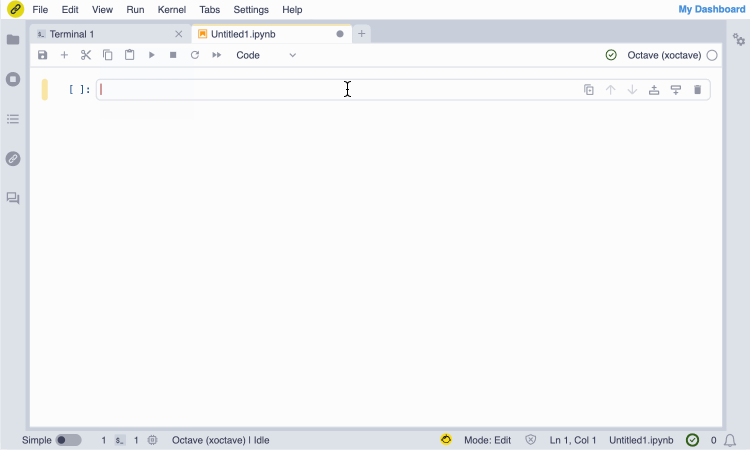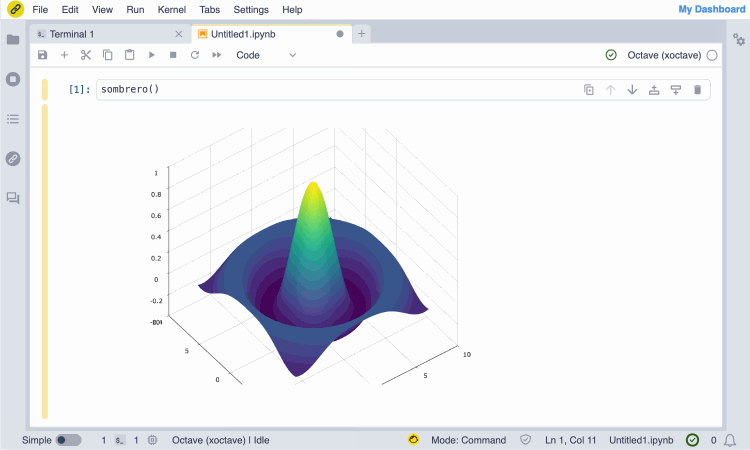 Screencast of the installation of the GNU Octave JupyterLite kernel from the notebook.link in-browser terminal, and immediate access with the notebook UI.
Screencast of the installation of the GNU Octave JupyterLite kernel from the notebook.link in-browser terminal, and immediate access with the notebook UI.
(The terminal session entirely runs in the browser and does not require any server.)
Beyond notebooks: AI-Assisted Workflows
Notebook.link does not only offer notebooks, but also integrates the JupyterLite-AI coding assistant.
The notebook.link chat UI enables rich MIME type rendering and in-browser tool calling, exposing all JupyterLab commands to the LLM. Expanding upon the core commands exposed by JupyterLab, we enabled the installation of WebAssembly conda packages by the LLM, enabling WebAssembly sandboxed workflows for exploratory data analysis, and removing the notebook entirely from the user workflow.
In the screencast below, we show how we can use Mistral AI models to install WebAssembly builds of Scikit-learn and Matplotlib, generate synthetic data, perform a logistic regression, and display a figure inline in the chat UI.
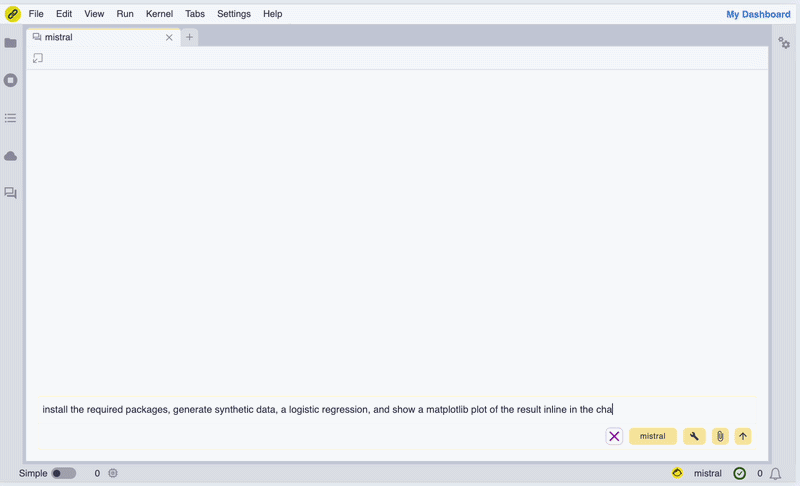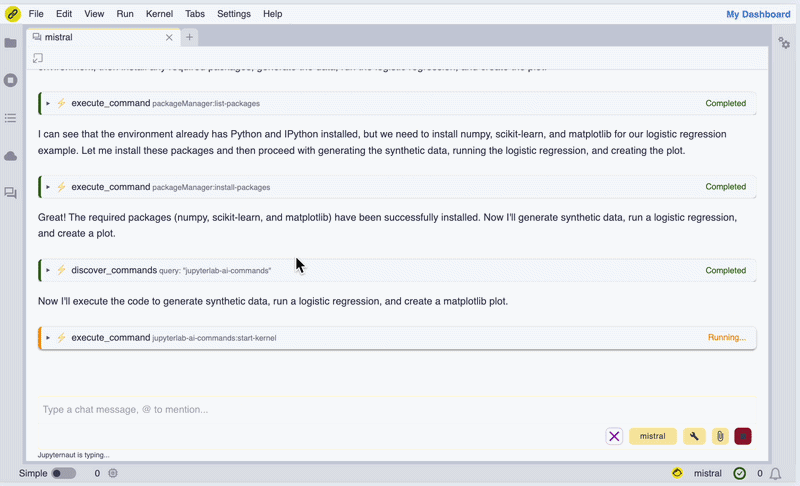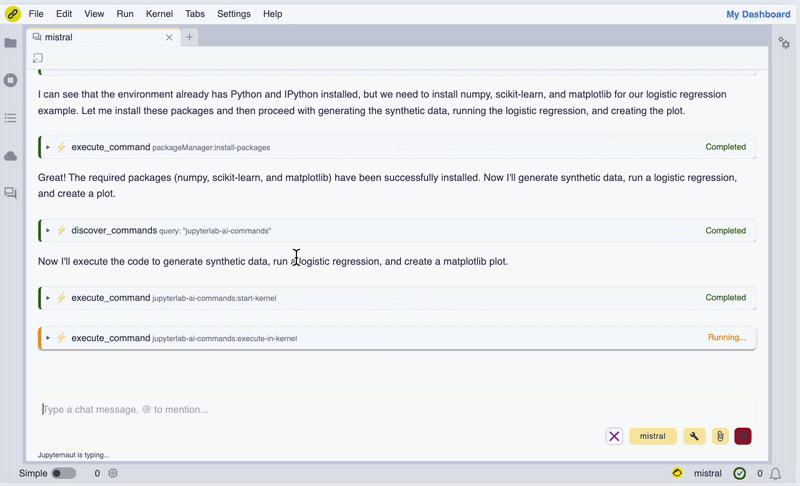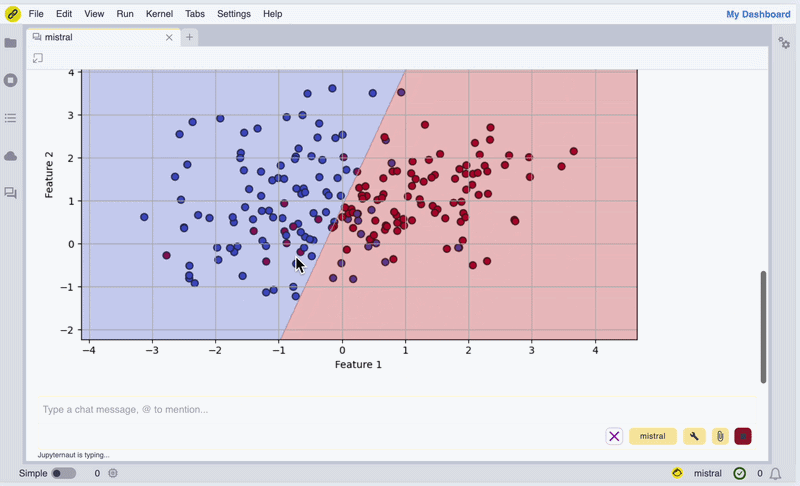 Screencast of the notebook.link AI assistant in action.
Screencast of the notebook.link AI assistant in action.
Conclusion: Scalable, Serverless Scientific Computing
Notebook.link represents a new model for scientific computing: open, scalable, and serverless. By leveraging JupyterLite, prefix.dev’s modern conda stack, and emscripten-forge, it delivers the full power of conda environments without traditional scaling constraints.
If you are interested in notebook.link for your organisation, integrated with your system - Single Sign-on (SSO), Learning Tools Interoperability (LTI) - then please contact us (QuantStack) for a quote and a deployment of notebook.link!
By Sylvain Corlay, on behalf of the Notebook.link team at QuantStack.
Thorsten Beier, Meriem Ben Ismail, Anutosh Bhat, Nicolas Brichet, Denisa Checiu, Afshin Darian, Florence Haudin, Yahia Heni, Trung Le, Johan Mabille, Isabel Paredes, Martin Renou, Anastasiia Sliusar, Ian Thomas, Andreas Trawöger, Jeremy Tuloup, Gabriela Vives